Advance/NeuralMD is a software for molecular dynamics calculations based on a Neural Network force field.
Using the results from first-principles calculations output by Quantum ESPRESSO as training data, you can create a molecular force field. This force field will then be employed to perform molecular dynamics simulations using LAMMPS. It incorporates various state-of-the-art technologies, such as automatic force field generation using self-learning hybrid Monte Carlo methods, acceleration on GPUs to enable calculations for systems with up to 100,000 atoms, and a pre-existing database of force fields.
Features of Advance/NeuralMD
(1) Force Field Definition Based on 3G-HDNNP
The Neural Network force field implemented in Advance/NeuralMD is based on the High-Dimensional Neural Network Potential (HDNNP). It also includes the utilization of a third-generation algorithm (3G-HDNNP) that accounts for long-range Coulomb interactions based on the charges of individual atoms. Furthermore, by combining our proprietary Δ-NNP method and techniques that utilize the average of multiple Neural Network models, we can create a stable force field with a relatively small amount of training data.
There are examples of applying these methods to calculate the lithium-ion conductivity of solid electrolytes and analyze the melting points of nuclear fuel materials.
Case 1: Conductivity Calculation of Lithium Ion Conductor LGPS by Δ-NNP Method
Case 2: Melting Point Analysis of Nuclear Fuel Materials Using Multiple Neural Network Models
(2) Operations from Advance/NanoLabo
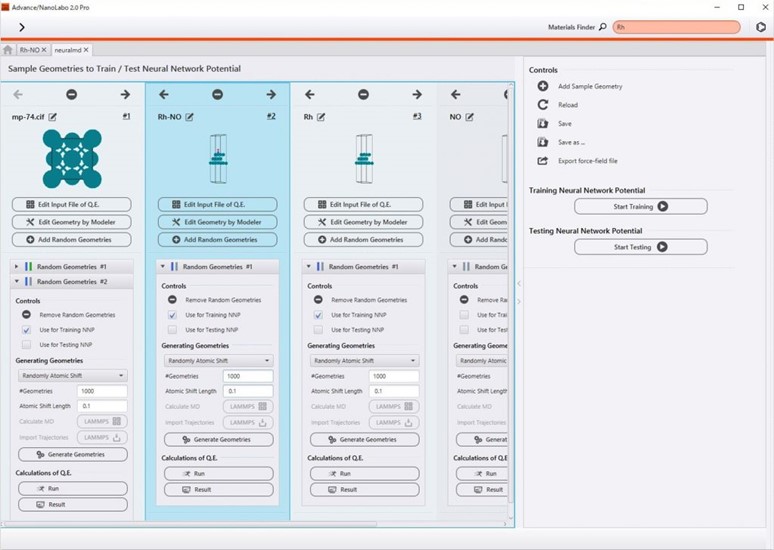
From creating training data, learning neural networks, and generating force fields to conducting molecular dynamics simulations, all processes can be operated from the Advance/NanoLabo interface. In the Grand Project (see the right figure), you can manage a large number of training data and proceed with your work. Furthermore, in the process of creating training data, it is also possible to distribute first-principles calculations to multiple computational resources such as calculation servers and the cloud.
(3) Self-Learning Hybrid Monte Carlo Method
Self-learning hybrid Monte Carlo method is an algorithm for first-principles Monte Carlo simulations developed by the Japan Atomic Energy Agency. By utilizing trajectories from molecular dynamics simulations with Neural Network force fields as proposed structures in the Monte Carlo method, it is possible to guarantee accuracy in Monte Carlo calculations comparable to first principles, while achieving efficient sampling of structures. Simultaneously with the execution of Monte Carlo calculations, the learning of the Neural Network force field is also conducted in parallel, using the results of first-principles calculations performed for each structure. As a result, when this method is executed, a Neural Network force field specialized for the target system is automatically generated.
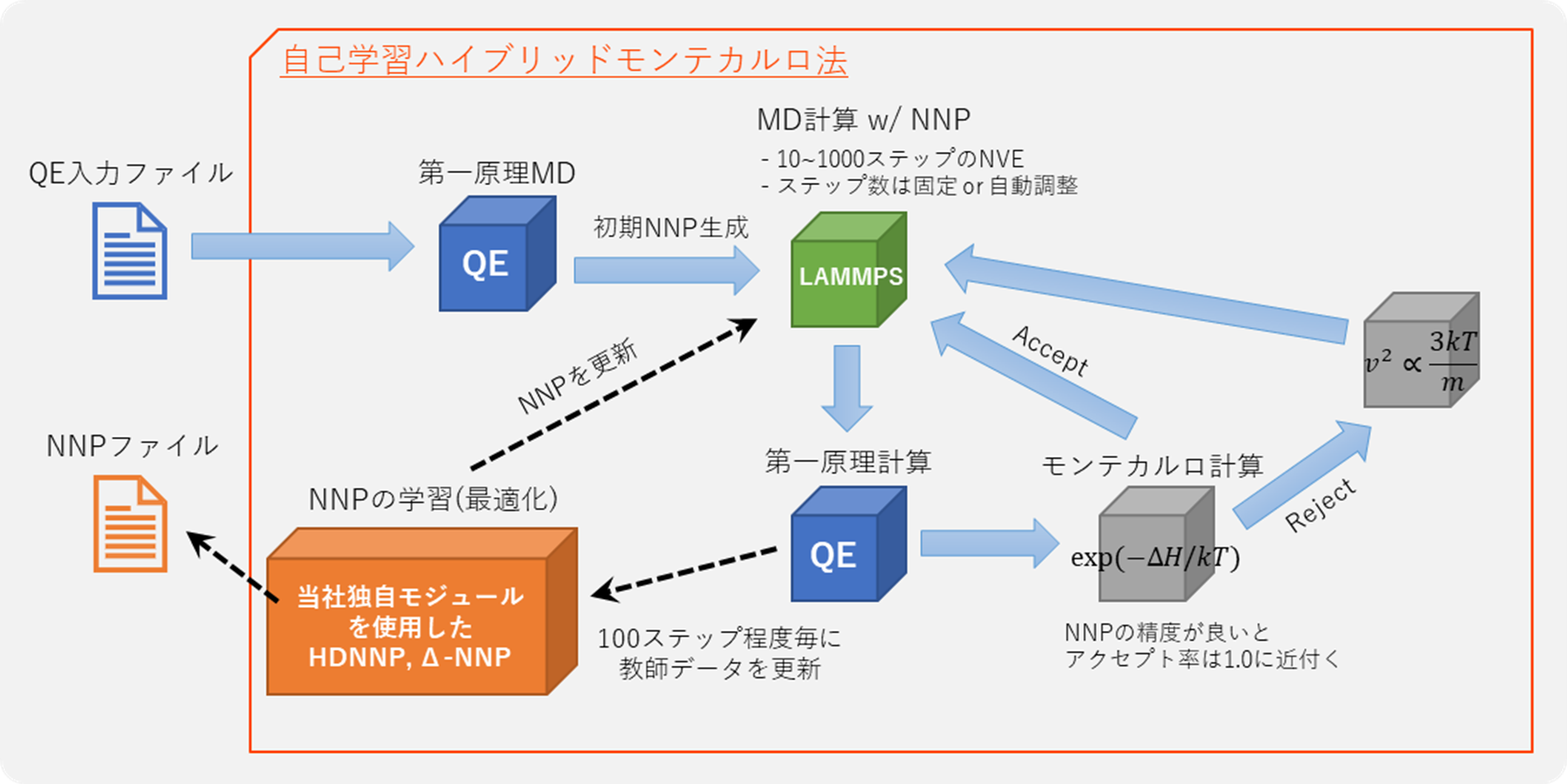
Case: Calculation of ion conductivity of oxide-based Lithium Ion Conductor LLZO via Self-Learning Hybrid Monte Carlo method
(4) Acceleration with GPU
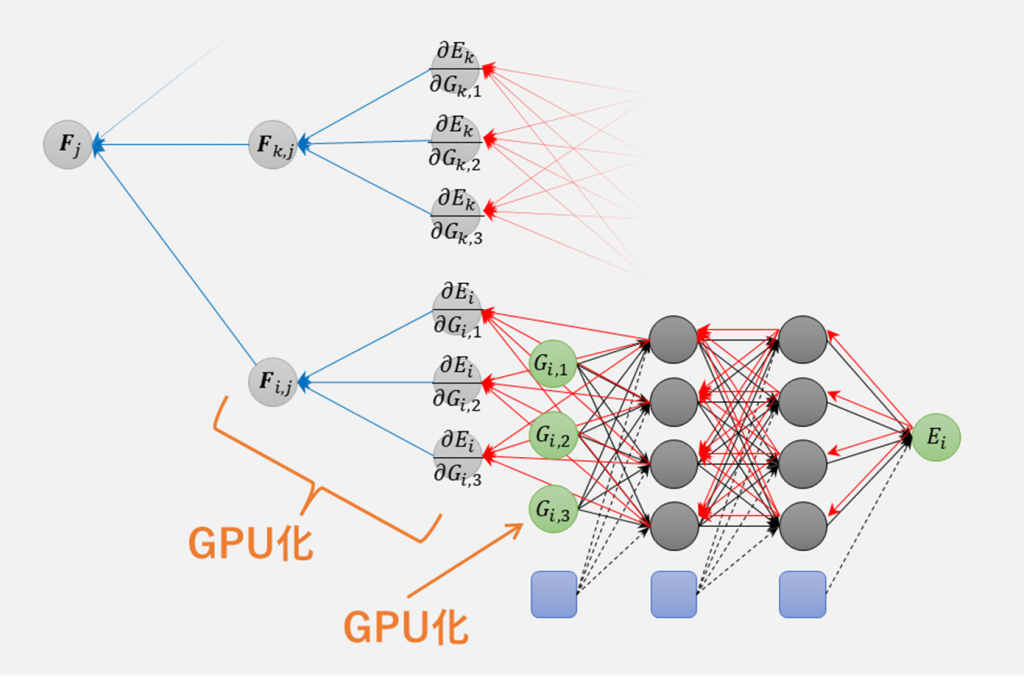
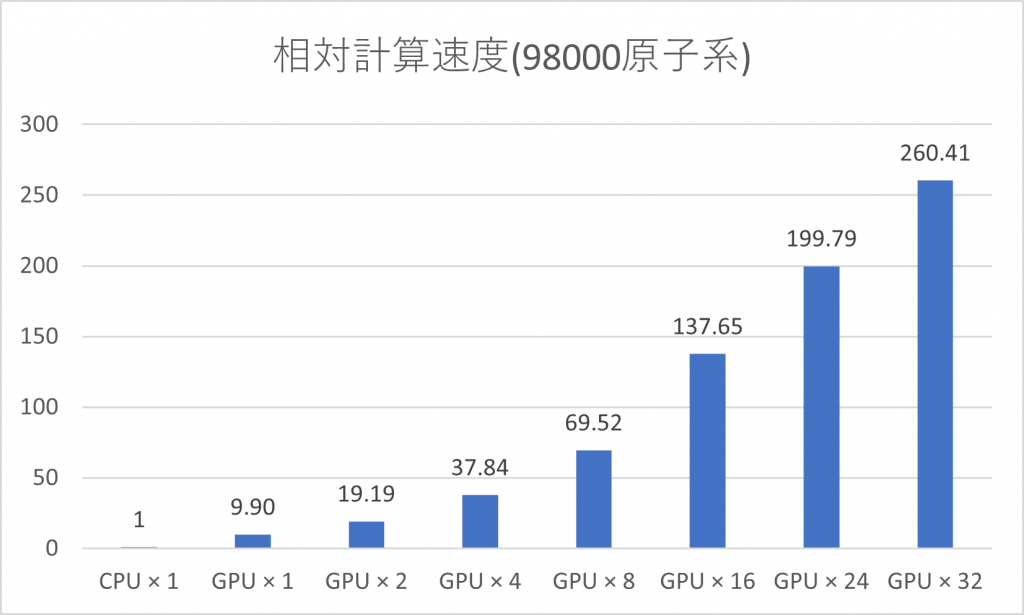
Advance/NeuralMD Pro supports neural network training and molecular dynamics simulations on GPUs. It can also be used in conjunction with MPI parallelization and is compatible with machines equipped with multiple GPUs and/or multiple machine nodes with GPUs. It is designed to maintain high utilization of both GPU and CPU by launching 2 to 4 MPI processes per GPU device. We have GPU-accelerated the computationally expensive symmetric function and force calculations (bottom left). By utilizing 32 GPUs, we have successfully achieved approximately 260 times faster molecular dynamics simulations for systems with around 100,000 atoms (using the computing environment provided by HPC systems).
Try Advance/NeuralMD (1 Month Free)